(We are once again very happy and grateful that Dr. Steffen Oeltze from the University of Magdeburg Visualization Group could write this short report on the medical visualization-related papers at IEEE VisWeek 2012 for us.)
This year, IEEE VisWeek hosted no special session dedicated to medical visualization but
five contributions to this field were spread over the conference program.
Jian Chen from the University of Maryland gave a talk on how stereo and screen size effect the legibility of three-dimensional streamtube visualizations. The effects were studied in the context of visually exploring dense fiber tracts reconstructed from diffusion magnetic resonance imaging data. A user study comprising 12 participants who had to perform five different tasks, e.g., find the endpoints of fiber tracts and judge if tracts belong to the same bundle, was carried out. In contrast to the initial hypotheses of Chen and
colleagues, completion time did not improve by using a larger display and performance accuracy was even hurt by introducing stereo. See the paper for further exploration of the results [1].
Rocco Gasteiger from the University of Magdeburg, Germany gave a compelling talk on the automatic detection and visualization of qualitative hemodynamic characteristics in cerebral aneurysms. He focused on the so-called inflow jet and the impingement zone, both being characteristics which are correlated with the risk of aneurysm rapture. Special care was taken in generating expressive visualizations of the detected features by means of glyphs, texture, Fresnel shading, and surface contours. The work was rounded off by a user study involving six domain experts who show high interpersonal variance in manually specifying the features but agreed on the good value of the presented visualizations [2]. A supplemental video can be watched here:
Markus Hadwiger from the King Abdullah University of Science and Technology, Saudi Arabia presented the first volume visualization system that scales to petascale volumes imaged as a continuous stream of high-resolution electron microscopy images. Markus and his team developed the system in collaboration with neuroscientists who wish to analyze brain function by measuring large blocks of brain tissue. The system can accept a constant stream of 2D image tiles from a microscope thereby handling missing data naturally and avoiding the expensive computation of any 3D multi-resolution representation such as an octree. A novelty of the system is that most computations are restricted to the currently visible volume data [3]. Watch the accompanying video here:
http://vimeo.com/50886921Alexander Bock from the Linköping University, Sweden presented the ray-casting of high-order finite element (FE) models for visualizing and analyzing strain of the human heart muscle. A straightforward ray-casting approach is inadequate for interactive data exploration due to the computational complexity of transforming the sample points along each ray into the non-uniform grid of the FE model. Hence, Alexander and his colleagues decoupled the expensive transformation from the rendering stage by means of proxy rays cast in FE space, thereby allowing it to be performed within a precomputation stage. The nice work was rounded off by an analysis of the error that is introduced by the presented approach [4].
Gunnar Läthén, also from the Linköping University, Sweden, gave a talk on improving transfer functions for volume rendering blood vessels in computed tomography angiography (CTA) data [5]. These data are acquired by means of injecting a contrast agent, which leads to an enhancement of the vessels. Due to variations in mixture concentration of contrast agent in the blood stream, the enhancement varies locally and transfer function presets often do not yield optimal images. Hence, Gunnar and his colleagues propose an automatic, optimization-based method that shifts transfer function presets to account for general deviations and local variations of the intensity of contrast enhanced blood vessels. The method is illustrated for clinically relevant CT angiography datasets.
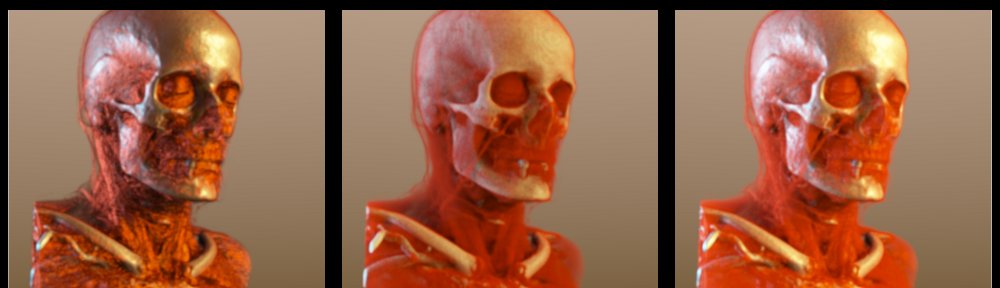
edited to add: I’ve received a tip from an anonymous reader that those interested in direct volume rendering might want to take a look at this work, which was also presented at VisWeek this year by Daniel Jönsson: Historygrams: Enabling Interactive Global Illumination in Direct Volume Rendering using Photon Mapping [6]

Example rendering from ‘Historygrams: Enabling Interactive Global Illumination in Direct Volume Rendering using Photon Mapping’.
References
- [1] Jian Chen; Haipeng Cai; Auchus, A.P.; Laidlaw, D.H.; , “Effects of Stereo and Screen Size on the Legibility of Three-Dimensional Streamtube Visualization,” IEEE Transactions on Visualization and Computer Graphics, vol.18, no.12, pp.2130-2139, Dec. 2012. URL: http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=6327218&isnumber=6327196
- [2] Gasteiger, R.; Lehmann, D.J.; van Pelt, R.; Janiga, G.; Beuing, O.; Vilanova, A.; Theisel, H.; Preim, B.; ,”Automatic Detection and Visualization of Qualitative Hemodynamic Characteristics in Cerebral Aneurysms,” IEEE Transactions on Visualization and Computer Graphics, vol.18, no.12, pp.2178-2187, Dec. 2012. URL: http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=6327222&isnumber=6327196
- [3] Hadwiger, M.; Beyer, J.; Won-Ki Jeong; Pfister, H.; , “Interactive Volume Exploration of Petascale Microscopy Data Streams Using a Visualization-Driven Virtual Memory Approach,” IEEE Transactions on Visualization and Computer Graphics, vol.18, no.12, pp.2285-2294, Dec. 2012. URL: http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=6327233&isnumber=6327196
- [4] Bock, A.; Sunden, E.; Bingchen Liu; Wunsche, B.; Ropinski, T.; , “Coherency-Based Curve Compression for High-Order Finite Element Model Visualization,” IEEE Transactions on Visualization and Computer Graphics, vol.18, no.12, pp.2315-2324, Dec. 2012. URL: http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=6327236&isnumber=6327196
- [5] Lathen, G.; Lindholm, S.; Lenz, R.; Persson, A.; Borga, M.; , “Automatic Tuning of Spatially Varying Transfer Functions for Blood Vessel Visualization,” IEEE Transactions on Visualization and Computer Graphics, vol.18, no.12, pp.2345-2354, Dec. 2012. URL: http://www.computer.org/csdl/trans/tg/2012/12/ttg2012122345-abs.html
- [6] Jönsson, D; Kronander, J.; Ropinski, T; Ynnerman, A.; , “Historygrams: Enabling Interactive Global Illumination in Direct Volume Rendering using Photon Mapping”, IEEE Transactions on Visualization and Computer Graphics, vol.18, no.12, pp.2364-2371, Dec. 2012. URL: http://www.computer.org/csdl/trans/tg/2012/12/ttg2012122364-abs.html